|
This structure is caused by chemical interactions between various amino acids and regions of the polypeptide. The unique three-dimensional structure of a polypeptide is known as its tertiary structure. Polymer Sci., A-1, 5, 2867 (1967).įinegold, L., and Cude, J. The -helix and -pleated sheet structures are found in many globular and fibrous proteins. V., New Problems in the Physics of Glass (OTS Department of Commerce, District of Columbia, 1963). Kittel, C., Introduction to Solid State Physics (Wiley, New York, 1971). Quantitatively, these measurements also provide tests of theories of published vibrational models of homopolypeptides, and will give hydrogen bond strengths. The low temperature measurements presented here are the first made on a polypeptide in alpha and beta-conformation, and clearly show the one-dimensional nature of the α-helix and the two-dimensional nature of the β-sheet structure. The important lower-energy modes are well examined by specific heat measurements, which do not require single crystal samples, at low temperatures (for example, 1–20 K, where only the acoustic modes are excited thermally to any appreciable extent-1 K ∼0.7 cm −1). Basically, however, these latter methods examine the optical vibrational modes of a system, that is, those modes which are of energy higher than, for example, 70 cm −1. Recent advances in high polymer and solid state physics have enabled the vibrational aspects of the simpler homopolypeptides to be treated as normal-but complicated-polymers by the theoretical techniques of lattice dynamics based on the experimental methods of neutron, infrared and Raman spectroscopy. Note that because of processes such as the post-translational modifications to proteins we still need protein sequencing and I believe that we currently rely too heavily on DNA sequencing.HOMOPOLYPEPTIDES provide good model systems for various aspects of proteins 1,2. 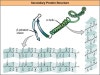
This is because it is now much easier to sequence DNA. Instead, since it has been worked out (mostly) how DNA codes for protein, we usually infer the protein sequence from the DNA sequence. 
In beta sheets amino acid chain is in an almost fully extended conformation, linear or ‘sheet like’. However, it is now relatively rare to directly determine protein sequence! Beta-pleated sheet (beta sheet) In alpha helix amino acid chain is in a right-handed spiral conformation or clockwise. It is the most widely common secondary protein. Alpha helix protein: Linus Pauling was the first scientist who observed the alpha helix structure by x-ray crystallography. alpha helix and beta pleated sheets of protein. 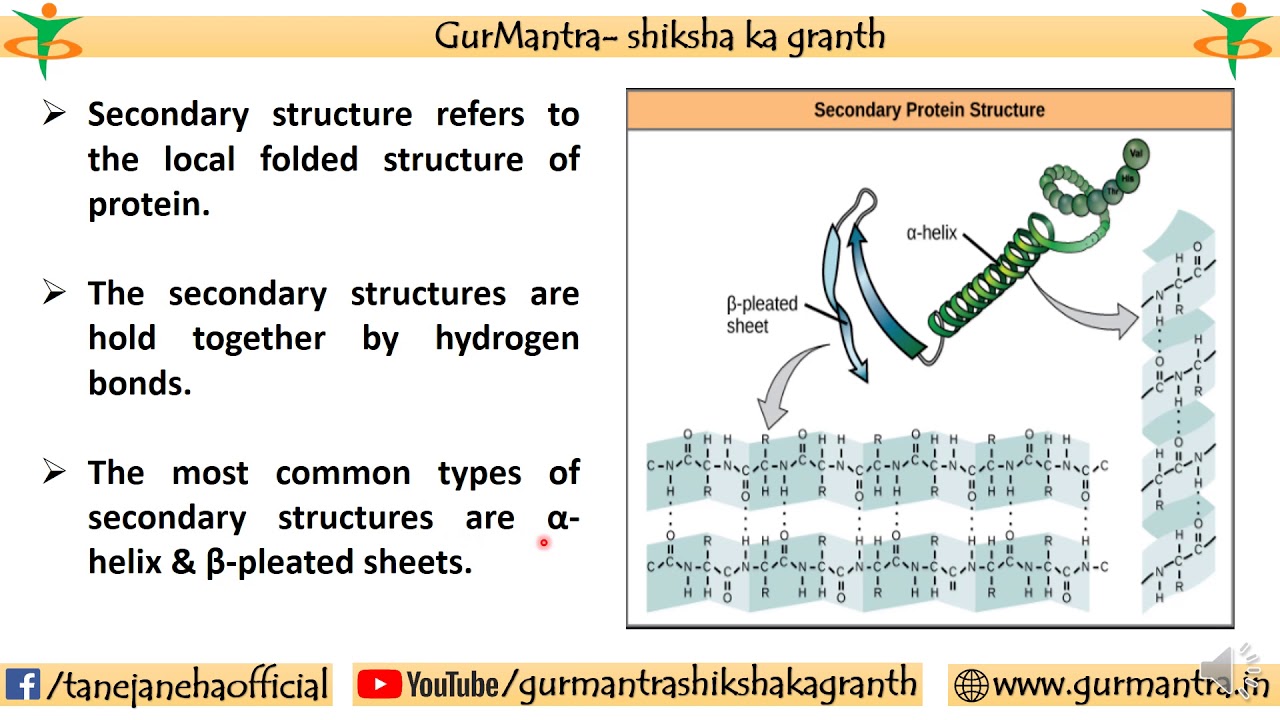
The most common type of secondary structure in proteins is the -helix. The secondary structure of a protein may be in two forms. The very first protein sequence (bovine insulin) was determined by Fredrick Sanger in 1951-2 (note that this was more than a decade before the first nucleotide sequence). Protein secondary structure: alpha-helices and beta-sheets, hairpins and loops. There are many different techniques for directly determining protein sequences - this wikipedia article is a decent introduction: There are also methods that have been developed to remove amino acids one at a time.īy combining theses techniques it is possible to directly determine protein sequences. This is a great question, but actually quite complicated so I'm not going to try to give a complete answer - I have given some useful links below if you wish to learn more.Įach amino acid has unique chemical properties that can be used to tell them apart.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
